Research Topics
Spatial omic imaging and machine learning are rapidly transforming the nature of biology research, providing rich avenues for discovery at the nexus of AI and mechanistic interpretation. My lab uses computational, mathematical, and high-throughput data generation approaches to study how cancer ecosystems function, evolve, and respond to therapeutic treatment. We study problems in cancer sequence and image analysis across a wide spectrum of cancer types, with particular expertise in breast cancer and patient-derived xenografts. These projects involve collaborations with experimental and computational colleagues at JAX Genomic Medicine, JAX Mammalian Genetics, and a number of outside groups.
For the most up-to-date information of the lab's work, see the Publications page.
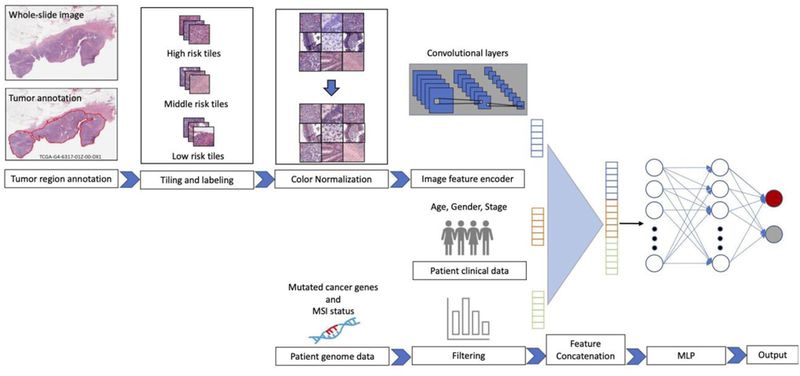
Computational Tissue Image AnalysisCancer treatment response is affected by tumor cell heterogeneity, immune cells, and other microenvironmental features, analogous to factors important in macroscopic ecosystems. Our lab uses advanced computational image analysis to reveal the spatial processes in tumors. We have developed convolutional neural networks that can be applied to H&E images to accurately distinguish tumor regions, identify cancer subtypes, and classify mutation status. Critically, our work has shown that images from diverse cancer types can be combined to enhance discovery from large image sets. The lab analyzes many types of imaging data, including spatial transcriptomics (Visium, Xenium, VisiumHD, merFISH), spatial proteomics (CODEX, imaging mass cytometry), 3D confocal microscopy, spatial long-read sequencing, and H&E/IHC across tumors, organoids, and xenografts. Our studies integrate discussions among computational biologists, cancer biologists, oncologists and pathologists.
Our lab co-leads the Data Analysis Core for the KAPP-Sen U54 that is part of the NIH Cellular Senescence Network (SenNet). In this project we perform spatial analysis of H&E, spatial transcriptomics, and spatial proteomics datasets for kidney, adipose, pancreas, and placental tissues, among others.
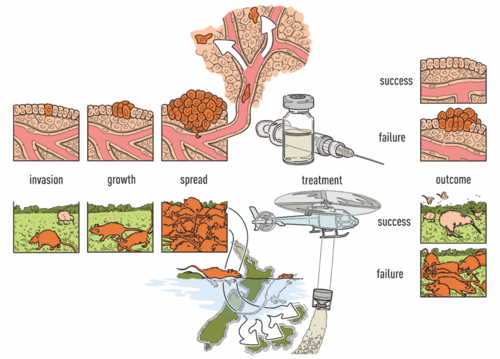
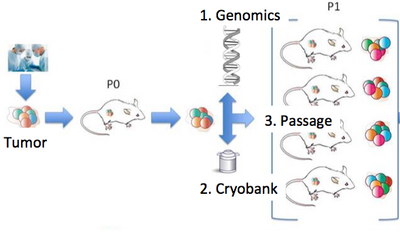
Our lab is a leader in the field of patient-derived xenografts, a model system in which human tumors are engrafted and studied in NSG mice. Xenografts play a critical role in cancer research, as they are often used in therapeutic testing just prior to clinical trials. JAX has developed >300 such models from cancer types including breast, lung, bladder, and others, and these are a community wide resource. Since 2017, our lab co-leads the Data Coordination Center for the NCI PDXNet, a multi-institute consortium supported by the NCI Cancer Moonshot Initiative. We are partnering with institutes around the country including the Huntsman Cancer Institute, Baylor College of Medicine, MD Anderson, Washington University, the Wistar Institute, NCI's Frederick National Laboratory, and Seven Bridges Genomics to jointly study cancer using xenografts. Our lab uses these to understand the genetic drivers of cancer and drug resistance, with a focus on computational modeling of intratumoral evolution (Woo, Giordano et al 2021, Kim et al 2018, Noorbakhsh and Chuang 2017) and also the robustness of xenografts for preclinical testing therapeutic agents (Evard et al 2020). For more information see the PDXNetwork site.
Within JAX we work closely with a number of groups studying cancer with xenografts and clinical samples, including the Liu, Bult, Lee, Palucka, Robson, and Anczukow-Camarda labs, on projects such as triple negative breast cancer (Menghi et al 2016), gastric cancer (Cho et al 2018), and immune cell activity (Yu et al, 2021).